Keywords
crystal structure
histone-like HU protein
NMR
nuclear magnetic resonance spectroscopy
SAXS
small-angle X-ray scattering
structure in solution
X-ray diffraction
XRD
Abstract
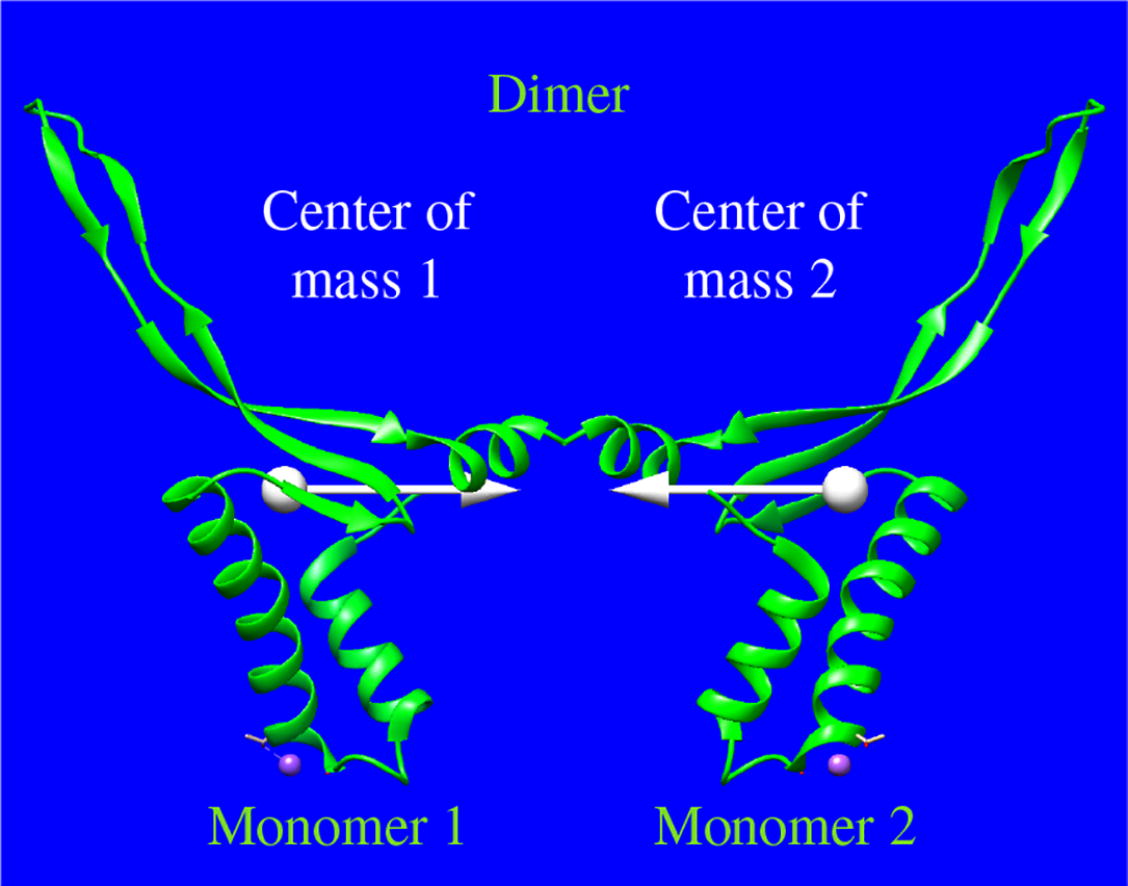
A solution of a histone-like protein from Spiroplasma melliferum (HUSpm) was examined by small-angle X-ray scattering (SAXS). The experimental SAXS curve was compared with those calculated for the HUSpm structures from the PDB databank obtained by both X-ray diffraction analysis and nuclear magnetic resonance spectroscopy. The model of the HUSpm structure in solution, which best agrees with the experimental SAXS data, has a shorter distance between the centers of mass of the HUSpm monomers compared to the crystal structure, indicating that the HUSpm monomers can be located closer to each other in solution than in the crystalline state.
References
1.

Keiffer S., Carneiro M.G., Hollander J., Kobayashi M., Pogoryelev D., AB E., Theisgen S., Müller G., Siegal G.
Journal of Biomolecular NMR,
2020
2.

Carlon A., Ravera E., Andrałojć W., Parigi G., Murshudov G.N., Luchinat C.
Progress in Nuclear Magnetic Resonance Spectroscopy,
2016
3.
10.1016/j.mencom.2022.11.011_b0015
Grove
Curr. Issues Mol. Biol.,
2011
4.

Kamashev D., Agapova Y., Rastorguev S., Talyzina A.A., Boyko K.M., Korzhenevskiy D.A., Vlaskina A., Vasilov R., Timofeev V.I., Rakitina T.V.
PLoS ONE,
2017
5.

Dame R.T., Rashid F.M., Grainger D.C.
Nature Reviews Genetics,
2019
6.

Stojkova P., Spidlova P., Stulik J.
Frontiers in Cellular and Infection Microbiology,
2019
7.

Hołówka J., Zakrzewska-Czerwińska J.
Frontiers in Microbiology,
2020
8.

Bhowmick T., Ghosh S., Dixit K., Ganesan V., Ramagopal U.A., Dey D., Sarma S.P., Ramakumar S., Nagaraja V.
Nature Communications,
2014
9.

Agapova Y.K., Altukhov D.A., Timofeev V.I., Stroylov V.S., Mityanov V.S., Korzhenevskiy D.A., Vlaskina A.V., Smirnova E.V., Bocharov E.V., Rakitina T.V.
Scientific Reports,
2020
10.

Rodríguez-Zamora P.
Progress in Biophysics and Molecular Biology,
2020
11.

Bengtsen T., Holm V.L., Kjølbye L.R., Midtgaard S.R., Johansen N.T., Tesei G., Bottaro S., Schiøtt B., Arleth L., Lindorff-Larsen K.
eLife,
2020
12.

Larsen A.H., Wang Y., Bottaro S., Grudinin S., Arleth L., Lindorff-Larsen K.
PLoS Computational Biology,
2020
13.

Boyko K.M., Rakitina T.V., Korzhenevskiy D.A., Vlaskina A.V., Agapova Y.K., Kamashev D.E., Kleymenov S.Y., Popov V.O.
Scientific Reports,
2016
14.

Blanchet C.E., Spilotros A., Schwemmer F., Graewert M.A., Kikhney A., Jeffries C.M., Franke D., Mark D., Zengerle R., Cipriani F., Fiedler S., Roessle M., Svergun D.I.
Journal of Applied Crystallography,
2015
15.

Manalastas-Cantos K., Konarev P.V., Hajizadeh N.R., Kikhney A.G., Petoukhov M.V., Molodenskiy D.S., Panjkovich A., Mertens H.D., Gruzinov A., Borges C., Jeffries C.M., Svergun D.I., Franke D.
Journal of Applied Crystallography,
2021
16.

Timofeev V.I., Altukhov D.A., Talyzina A.A., Agapova Y.K., Vlaskina A.V., Korzhenevskiy D.A., Kleymenov S.Y., Bocharov E.V., Rakitina T.V.
Journal of Biomolecular Structure and Dynamics,
2018
17.

Pettersen E.F., Goddard T.D., Huang C.C., Couch G.S., Greenblatt D.M., Meng E.C., Ferrin T.E.
Journal of Computational Chemistry,
2004
18.
10.1016/j.mencom.2022.11.011_b0090
Persson
J. Chem. Phys.,
2018
19.

Sheu S., Yang D.
Journal of Physical Chemistry B,
2010
20.
L. Schrödinger and W. DeLano, PyMOL, 2020, https://www.pymol.org/ pymol.html.