Keywords
N-acetyl-l-aspartate
aspartate N-acetyltransferase
Gibbs energy profiles
molecular dynamics
NAT8L
QM/MM potentials
Reaction mechanism
Abstract
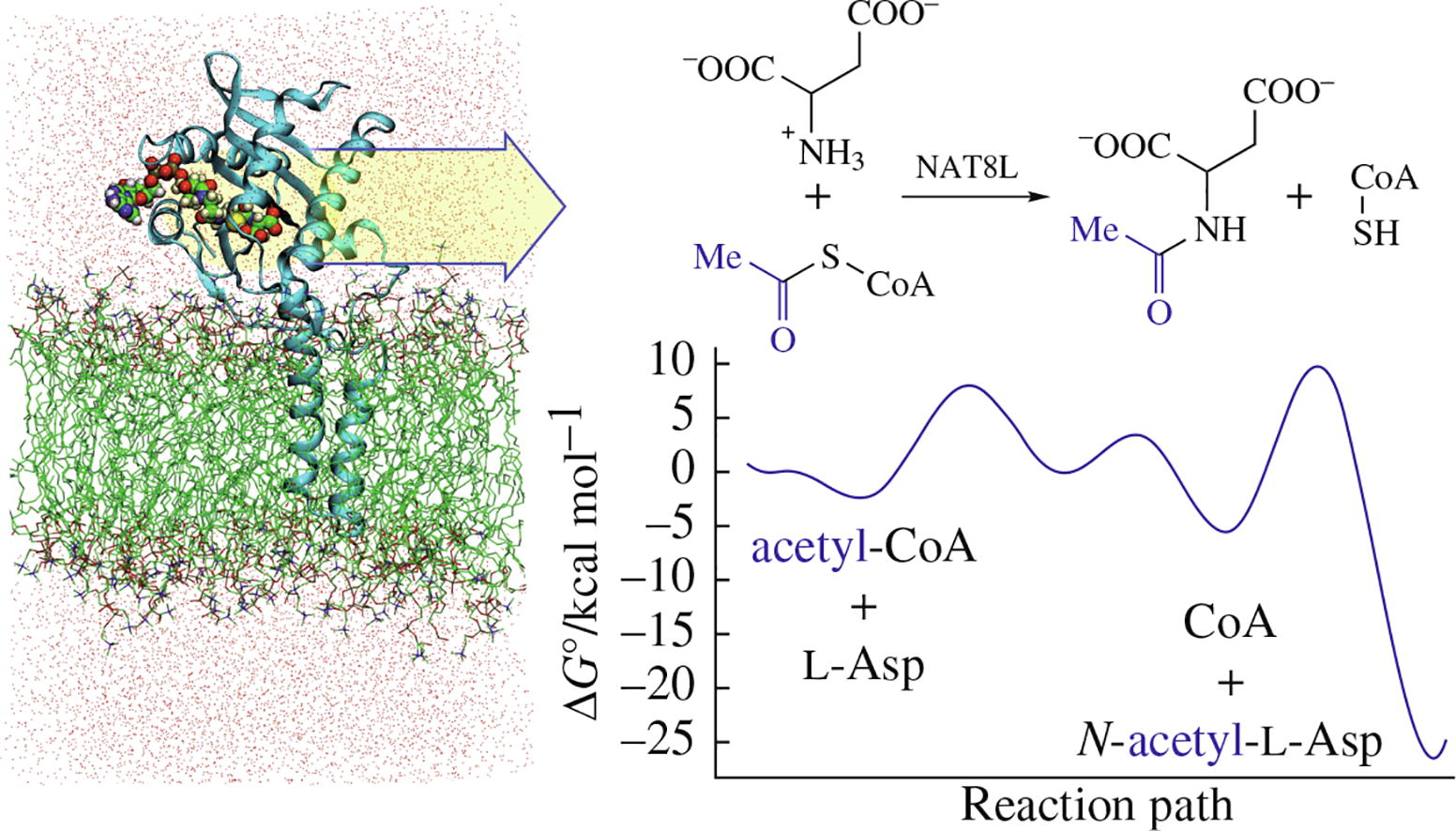
The results of a computational study of the synthesis of a key brain metabolite, N-acetyl-l-aspartate, catalyzed by aspartate N-acetyltransferase, encoded by the NAT8L gene, are reported. The reaction Gibbs energy profiles were computed using molecular dynamics simulations with interaction potentials estimated on-the-fly by the quantum mechanics/molecular mechanics QM(PBE0/6-31G**)/MM(CHARMM) approach. The revealed reaction mechanism includes four elementary steps with corresponding activation energies not exceeding 14 kcal mol−1
References
1.

MOFFETT J., ROSS B., ARUN P., MADHAVARAO C., NAMBOODIRI A.
Progress in Neurobiology,
2007
2.

Ariyannur P.S., Moffett J.R., Manickam P., Pattabiraman N., Arun P., Nitta A., Nabeshima T., Madhavarao C.N., Namboodiri A.M.
Brain Research,
2010
3.

Hussain M.S., Wang Q., Viola R.E.
Archives of Biochemistry and Biophysics,
2021
4.

Dyda F., Klein D.C., Hickman A.B.
Annual Review of Biophysics and Biomolecular Structure,
2000
5.

Tahay G., Wiame E., Tyteca D., Courtoy P., Van Schaftingen E.
Biochemical Journal,
2011
6.
10.1016/j.mencom.2022.11.010_b0030
Polyakov
Neurosci.,
2020
7.
10.1016/j.mencom.2022.11.010_b0035
Varfolomeev
Neurosci.,
2020
8.

Varfolomeev S.D., Bykov V.I., Tsybenova S.B.
Doklady Biochemistry and Biophysics,
2020
9.

Dall'Acqua W., Carter P.
Protein Science,
2008
10.

Melo M.C., Bernardi R.C., Rudack T., Scheurer M., Riplinger C., Phillips J.C., Maia J.D., Rocha G.B., Ribeiro J.V., Stone J.E., Neese F., Schulten K., Luthey-Schulten Z.
Nature Methods,
2018
11.
10.1016/j.mencom.2022.11.010_b0055
Khrenova
Khim. Fiz.,
2022
12.

Phillips J.C., Hardy D.J., Maia J.D., Stone J.E., Ribeiro J.V., Bernardi R.C., Buch R., Fiorin G., Hénin J., Jiang W., McGreevy R., Melo M.C., Radak B.K., Skeel R.D., Singharoy A., et. al.
Journal of Chemical Physics,
2020
13.

Phillips J.C., Braun R., Wang W., Gumbart J., Tajkhorshid E., Villa E., Chipot C., Skeel R.D., Kalé L., Schulten K.
Journal of Computational Chemistry,
2005
14.

Seritan S., Bannwarth C., Fales B.S., Hohenstein E.G., Isborn C.M., Kokkila‐Schumacher S.I., Li X., Liu F., Luehr N., Snyder J.W., Song C., Titov A.V., Ufimtsev I.S., Wang L., Martínez T.J., et. al.
Wiley Interdisciplinary Reviews: Computational Molecular Science,
2020
15.

Khrenova M.G., Tsirelson V.G., Nemukhin A.V.
Physical Chemistry Chemical Physics,
2020
16.

Best R.B., Zhu X., Shim J., Lopes P.E., Mittal J., Feig M., MacKerell A.D.
Journal of Chemical Theory and Computation,
2012
17.
10.1016/j.mencom.2022.11.010_b0085
Denning
J. Comput. Chem.,
1929
18.

Jorgensen W.L., Chandrasekhar J., Madura J.D., Impey R.W., Klein M.L.
Journal of Chemical Physics,
1983
19.

Vanommeslaeghe K., MacKerell A.D.
Journal of Chemical Information and Modeling,
2012
20.

Vanommeslaeghe K., Raman E.P., MacKerell A.D.
Journal of Chemical Information and Modeling,
2012
21.
10.1016/j.mencom.2022.11.010_b0105
Kästner
Wiley Interdiscip. Rev.: Comput. Mol. Sci.,
2011
22.
10.1016/j.mencom.2022.11.010_b0110
Kästner
J. Chem. Phys.,
2005