Keywords
allosteric site of AMPA receptor
cycloprolylglycine
endogenous ampakine
MM-GBSA
piracetam
Abstract
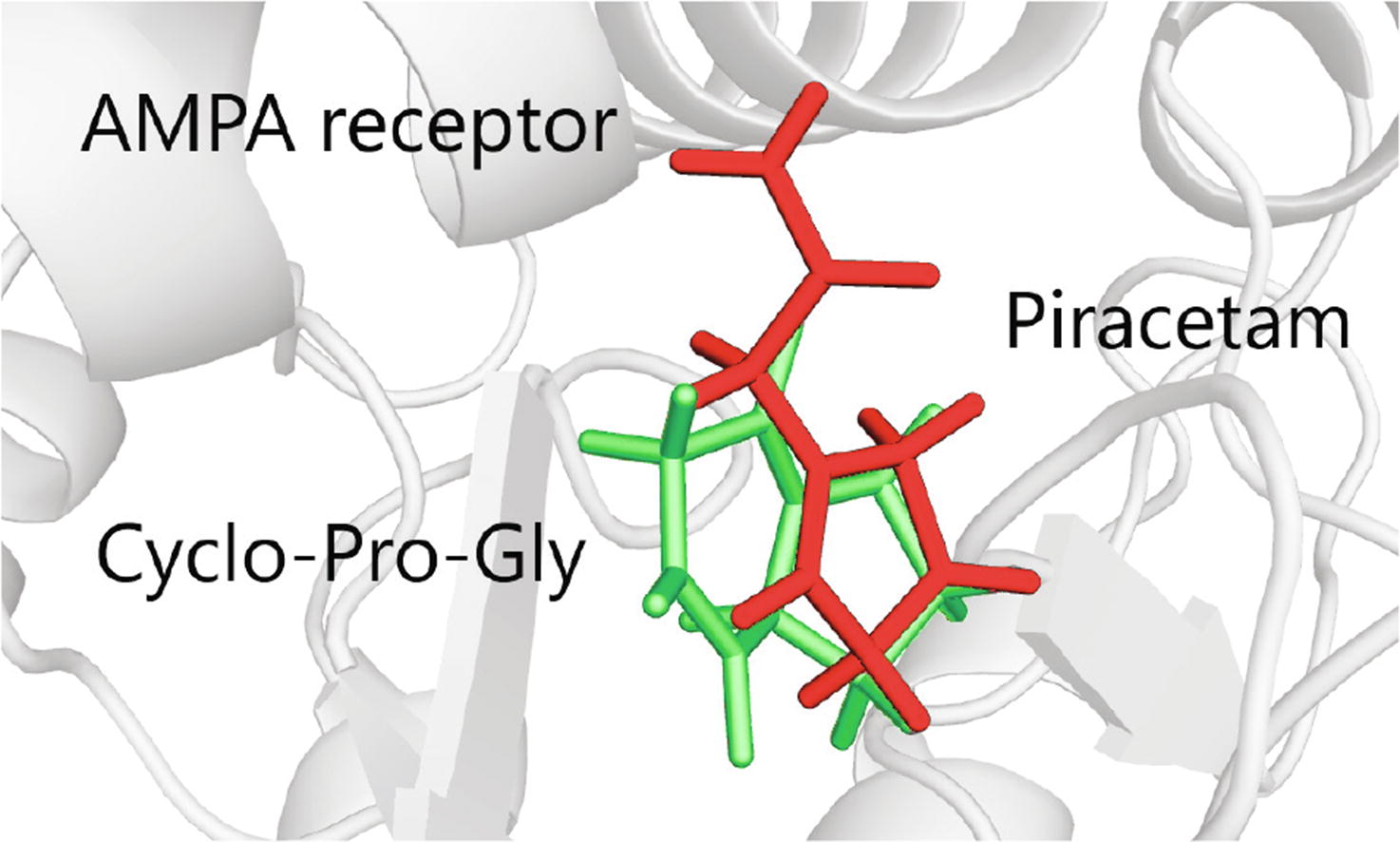
The calculations using the MM-GBSA method demonstrated that the neuropeptide cycloprolylglycine, structurally and pharmacologically resembling the classical nootropic piracetam, can bind to the piracetam site of the ligand- binding domain of the GluA3i AMPA receptor subtype with the binding energy higher than that of piracetam. Thus, together with the previous electrophysiological and biochemical data, the computational results confirm that cycloprolylglycine may be an endogenous ampakine.
References
1.

Gudasheva T.A., Boyko S.S., Akparov V.K., Ostrovskaya R.U., Skoldinov S.P., Rozantsev G.G., Voronina T.A., Zherdev V.P., Seredenin S.B.
FEBS Letters,
1996
2.

Fan D., Krishnamurthi R., Harris P., Barber P.A., Guan J.
Annals of Clinical and Translational Neurology,
2019
3.

Gudasheva T.A., Ostrovskaya R.U., Trofimov S.S., Voronina T.A., Skoldinov A.P., Seredenin S.B.
Bulletin of Experimental Biology and Medicine,
1999
4.
10.1016/j.mencom.2022.07.003_b0020
Koliasnikova
Eksp. Klin. Farmakol.,
2012
5.

Povarnina P.Y., Kolyasnikova K.N., Nikolaev S.V., Antipova T.A., Gudasheva T.A.
Bulletin of Experimental Biology and Medicine,
2016
6.

Gudasheva T.A., Konstantinopol'skii M.A., Ostrovskaya R.U., Seredenin S.B.
Bulletin of Experimental Biology and Medicine,
2001
7.

Gudasheva T.A., Grigoriev V.V., Koliasnikova K.N., Zamoyski V.L., Seredenin S.B.
Doklady Biochemistry and Biophysics,
2016
8.

Gudasheva T.A., Povarnina P.Y., Koliasnikova K.N., Alyaeva A.G., Vorontsova O.N., Seredenin S.B.
Doklady Biochemistry and Biophysics,
2020
9.

Copani A., Genazzani A.A., Aleppo G., Casabona G., Canonico P.L., Scapagnini U., Nicoletti F.
Journal of Neurochemistry,
1992
10.

Ahmed A.H., Oswald R.E.
Journal of Medicinal Chemistry,
2010
11.

Ptak C.P., Ahmed A.H., Oswald R.E.
Biochemistry,
2009
12.
Temnyakova N.S., Vasilenko D.A., Lavrov M.I., Karlov D.S., Grishin Y.K., Zamoyski V.L., Grigoriev V.V., Averina E.B., Palyulin V.A.
Mendeleev Communications,
2021
13.
Temnyakova N.S., Vasilenko D.A., Lavrov M.I., Karlov D.S., Grishin Y.K., Zamoyski V.L., Grigoriev V.V., Averina E.B., Palyulin V.A.
Mendeleev Communications,
2021
14.

Kollman P.A., Massova I., Reyes C., Kuhn B., Huo S., Chong L., Lee M., Lee T., Duan Y., Wang W., Donini O., Cieplak P., Srinivasan J., Case D.A., Cheatham T.E., et. al.
Accounts of Chemical Research,
2000
15.

Srinivasan J., Trevathan M.W., Beroza P., Case D.A.
Theoretical Chemistry Accounts,
1999
16.

Emsley P., Lohkamp B., Scott W.G., Cowtan K.
Acta Crystallographica Section D Biological Crystallography,
2010
17.

Case D.A., Cheatham T.E., Darden T., Gohlke H., Luo R., Merz K.M., Onufriev A., Simmerling C., Wang B., Woods R.J.
Journal of Computational Chemistry,
2005
18.

Salomon-Ferrer R., Götz A.W., Poole D., Le Grand S., Walker R.C.
Journal of Chemical Theory and Computation,
2013
19.

Maier J.A., Martinez C., Kasavajhala K., Wickstrom L., Hauser K.E., Simmerling C.
Journal of Chemical Theory and Computation,
2015
20.
10.1016/j.mencom.2022.07.003_b0100
Allen
Computer Simulation of Liquids,
1991
21.

Berendsen H.J., Postma J.P., van Gunsteren W.F., DiNola A., Haak J.R.
Journal of Chemical Physics,
1984
22.

Darden T., York D., Pedersen L.
Journal of Chemical Physics,
1993
23.

Miller B.R., McGee T.D., Swails J.M., Homeyer N., Gohlke H., Roitberg A.E.
Journal of Chemical Theory and Computation,
2012
24.

Onufriev A., Bashford D., Case D.A.
Proteins: Structure, Function and Genetics,
2004