Keywords
Abstract
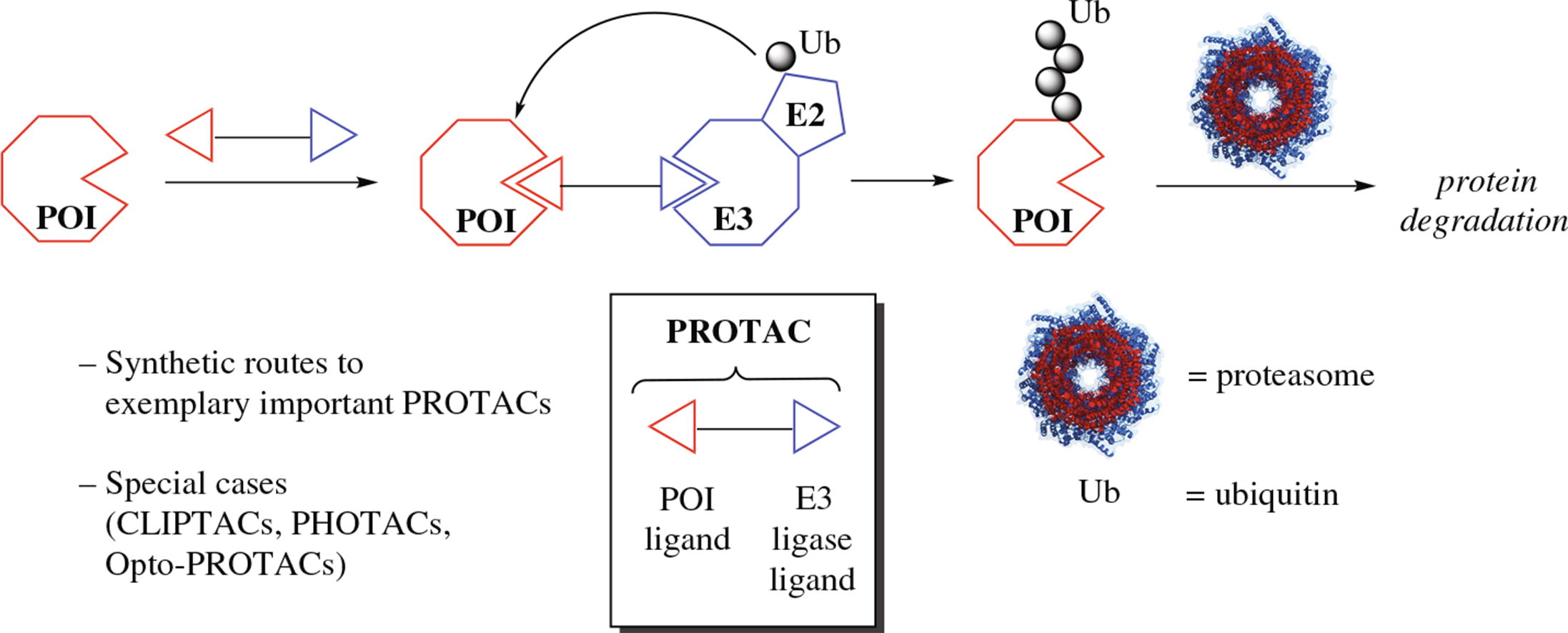
The development of various heterobifunctional constructs dubbed PRoteolysis-TArgeting Chimeras (PROTACs) has gained a significant impetus in the last few years. A viable alternative to the traditional occupancy-based inhibition of aberrantly hyperactive proteins, PROTACs operate by an event-based catalytic mechanism bringing together the protein of interest (POI, to be degraded) and E3 ubiquitin ligases. The formation of the ternary complex ‘POI–PROTAC–E3 ubiquitin ligase’ is the critical step which leads to the ubiquitination of the POI and its proteasomal degradation. The current Focused Review aims to highlight the syntheses of selected innovative PROTAC-type degraders of the therapeutically important protein targets as well as some notable chemical aspects of PROTAC construction. The overview is focusing on PROTACs aimed at recruiting Cereblon, the most exploited E3 ligase for targeted protein degradation.
References

























































![Discovery of 4-((3′R,4′S,5′R)-6″-Chloro-4′-(3-chloro-2-fluorophenyl)-1′-ethyl-2″-oxodispiro[cyclohexane-1,2′-pyrrolidine-3′,3″-indoline]-5′-carboxamido)bicyclo[2.2.2]octane-1-carboxylic Acid (AA-115/APG-115): A Potent and Orally Active Murine Double Minute 2 (MDM2) Inhibitor in Clinical Development](/storage/images/resized/iLiQsFqFaSEx6chlGQ5fbAwF6VYU3WWa08hkss0g_small_thumb.webp)