Keywords
COVID-19
molecular docking
molecular dynamics
natural compounds
SARS-CoV-2 inhibitors
SARS-CoV-2 Mpro
Abstract
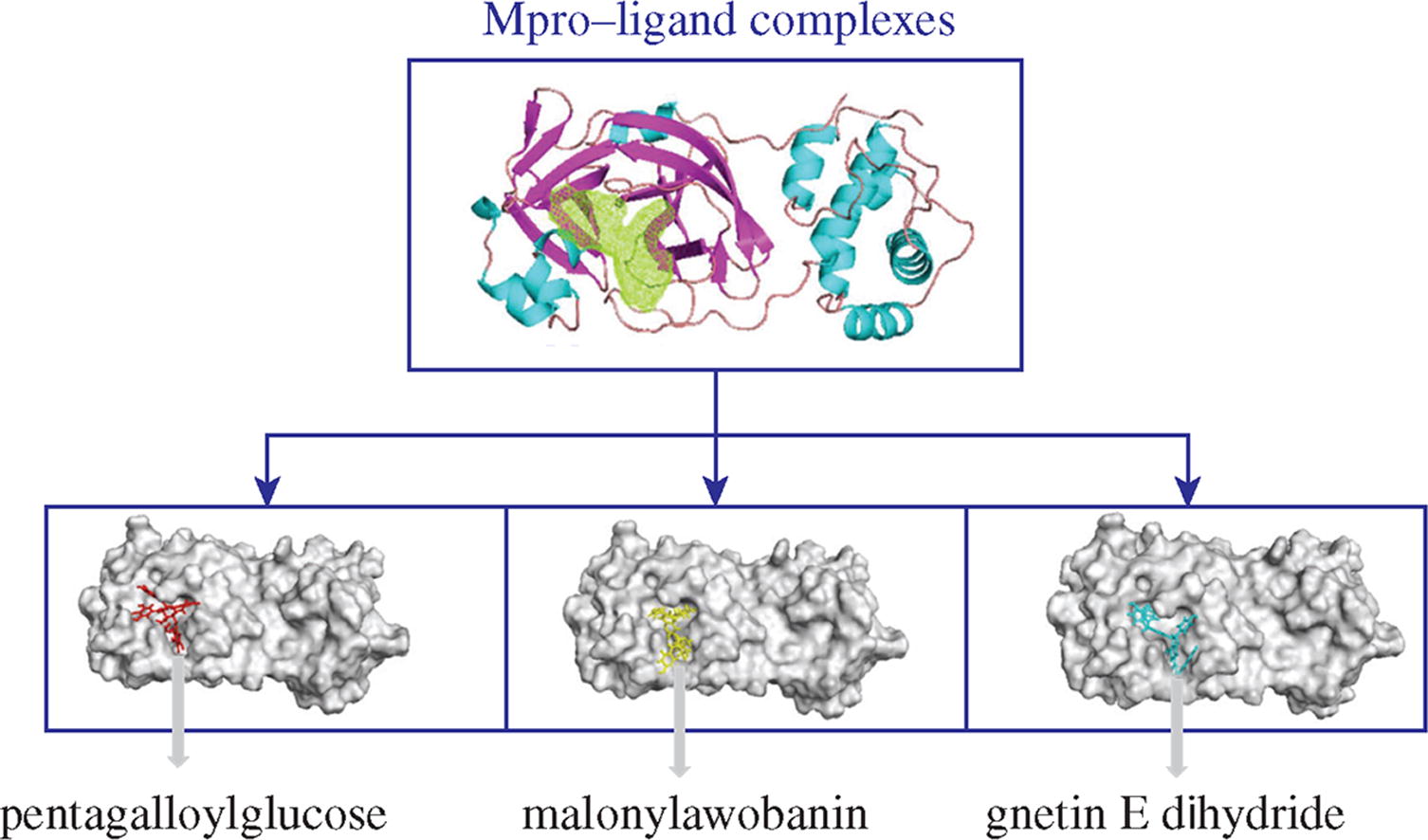
The SARS-CoV-2 main protease (Mpro) has been chosen as a conserved molecular target to develop broad-spectrum antiviral drugs. Using molecular docking and molecular dynamics (MD) simulations, a total of 5600 natural compounds available for virtual screening were tested to identify potential inhibitors of SARS-CoV-2 Mpro. As a result, three natural compounds (pentagalloylglucose, malonylawobanin and gnetin E dihydride) were found to be potential inhibitors of SARS-CoV-2, which confirms the theoretical and practical significance of this approach for the design of SARS-CoV-2 inhibitors.
References
1.

Chen J.
Microbes and Infection,
2020
2.

Cui J., Li F., Shi Z.
Nature Reviews Microbiology,
2018
3.

de Wit E., van Doremalen N., Falzarano D., Munster V.J.
Nature Reviews Microbiology,
2016
4.
WHO Coronavirus (COVID-19) Dashboard, World Health Organization, February 18, 2022. https://covid19.who.int/.
5.

Suleiman M.R., Wang H., Huang D., Wang H., Joseph J., Huang T., Zhang F., Wang J., Cheng M.
Journal of Biomolecular Structure and Dynamics,
2020
6.
Stroylov V.S., Svitanko I.V.
Mendeleev Communications,
2020
7.

Richardson P., Griffin I., Tucker C., Smith D., Oechsle O., Phelan A., Rawling M., Savory E., Stebbing J.
The Lancet,
2020
8.

Teralı K., Baddal B., Gülcan H.O.
Journal of Molecular Graphics and Modelling,
2020
9.

Elmezayen A.D., Al-Obaidi A., Şahin A.T., Yelekçi K.
Journal of Biomolecular Structure and Dynamics,
2020
10.

Gao Y., Yan L., Huang Y., Liu F., Zhao Y., Cao L., Wang T., Sun Q., Ming Z., Zhang L., Ge J., Zheng L., Zhang Y., Wang H., Zhu Y., et. al.
Science,
2020
11.

Subissi L., Posthuma C.C., Collet A., Zevenhoven-Dobbe J.C., Gorbalenya A.E., Decroly E., Snijder E.J., Canard B., Imbert I.
Proceedings of the National Academy of Sciences of the United States of America,
2014
12.

Zhou J., Fang L., Yang Z., Xu S., Lv M., Sun Z., Chen J., Wang D., Gao J., Xiao S.
FASEB Journal,
2019
13.

Wang F., Chen C., Tan W., Yang K., Yang H.
Scientific Reports,
2016
14.
15.

Wang M., Li W., Wang Y., Song Y., Wang J., Cheng M.
Journal of Molecular Graphics and Modelling,
2018
16.

Nguyen H.L., Thai N.Q., Truong D.T., Li M.S.
Journal of Physical Chemistry B,
2020
17.
Costanzo M., De Giglio M.A., Roviello G.N.
Current Medicinal Chemistry,
2020
18.

Jin Z., Du X., Xu Y., Deng Y., Liu M., Zhao Y., Zhang B., Li X., Zhang L., Peng C., Duan Y., Yu J., Wang L., Yang K., Liu F., et. al.
Nature,
2020
19.

Jia W., Jing Z., Liu X., Feng X., Liu Y., Wang S., Xu W., Liu J., Cheng X.
Journal of Biomolecular Structure and Dynamics,
2017
20.

Liu Y., Feng X., Jia W., Jing Z., Xu W., Cheng X.
Computational Biology and Chemistry,
2019
21.

Wang Y., Hu B., Peng Y., Xiong X., Jing W., Wang J., Gao H.
Journal of Chemical Information and Modeling,
2019
22.

Wang Z., Ma C., Yang J., Gao Q., Sun X., Ding L., Liu H.
Journal of Molecular Structure,
2019