Keywords
inhibition
molecular mechanism
QM/MM
thiocyanate dehydrogenase
thiourea
urea
Abstract
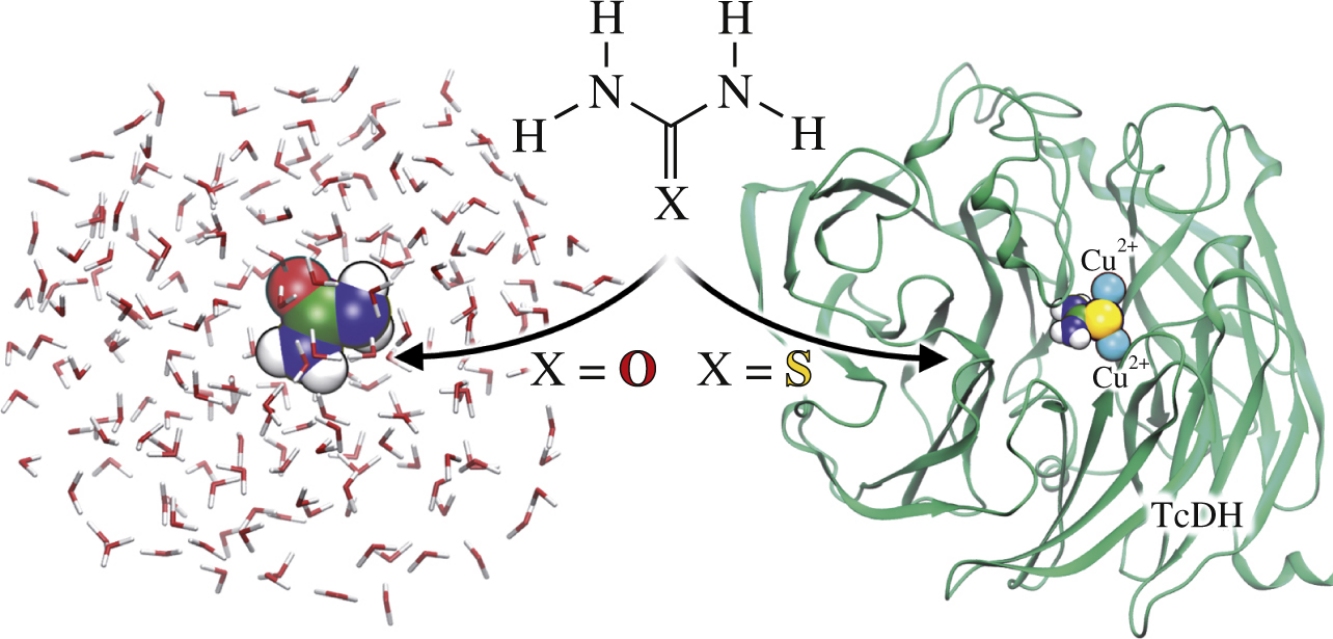
According to steady-state kinetic experiments, thiourea inhibits thiocyanate dehydrogenase TcDH, whereas urea does not. The QM/MM modeling combined with electron density analysis reveals the molecular mechanism of this process. For both compounds, interactions with bulk water molecules are similar, but upon binding to the active site of thiocyanate dehydrogenase a sulfur atom forms stronger coordination bonds with copper ions than an oxygen.
References
1.

Tikhonova T.V., Sorokin D.Y., Hagen W.R., Khrenova M.G., Muyzer G., Rakitina T.V., Shabalin I.G., Trofimov A.A., Tsallagov S.I., Popov V.O.
Proceedings of the National Academy of Sciences of the United States of America,
2020
2.

Stojceva Radovanovic B.C., Premovic P.I.
Journal of Thermal Analysis and Calorimetry,
1992
3.

Esmaeili L., Perez M.G., Jafari M., Paquin J., Ispas-Szabo P., Pop V., Andruh M., Byers J., Mateescu M.A.
Journal of Inorganic Biochemistry,
2019
4.

Razak N.H., Tan L.L., Hasbullah S.A., Heng L.Y.
Microchemical Journal,
2020
5.
10.1016/j.mencom.2021.04.030_bib0025
Bader
Atoms in Molecules – A Quantum Theory,
1990
6.
Markov A.A., Yakushev I.A., Churakov A.V., Khrustalev V.N., Cherkashina N.V., Stolarov I.P., Gekhman A.E., Vargaftik M.N.
Mendeleev Communications,
2019
7.
Vatsadze S.Z., Medved’ko A.V., Bodunov A.A., Lyssenko K.A.
Mendeleev Communications,
2020
8.

Khrenova M.G., Krivitskaya A.V., Tsirelson V.G.
New Journal of Chemistry,
2019
9.
Khrenova M.G., Tsirelson V.G.
Mendeleev Communications,
2019
10.
Khrenova M.G., Nemukhin A.V., Tsirelson V.G.
Mendeleev Communications,
2020
11.
Kulakova A.M., Khrenova M.G.
Mendeleev Communications,
2021
12.

Espinosa E., Molins E., Lecomte C.
Chemical Physics Letters,
1998
13.

Adamo C., Barone V.
Journal of Chemical Physics,
1999
14.

A Second Generation Force Field for the Simulation of Proteins, Nucleic Acids, and Organic Molecules
Cornell W.D., Cieplak P., Bayly C.I., Gould I.R., Merz K.M., Ferguson D.M., Spellmeyer D.C., Fox T., Caldwell J.W., Kollman P.A.
Journal of the American Chemical Society,
1995
15.

Jorgensen W.L., Chandrasekhar J., Madura J.D., Impey R.W., Klein M.L.
Journal of Chemical Physics,
1983
16.

Valiev M., Bylaska E.J., Govind N., Kowalski K., Straatsma T.P., Van Dam H.J., Wang D., Nieplocha J., Apra E., Windus T.L., de Jong W.A.
Computer Physics Communications,
2010
17.

Lu T., Chen F.
Journal of Computational Chemistry,
2011
18.
10.1016/j.mencom.2021.04.030_bib0090
Voevodin
Supercomputing Frontiers and Innovations,
2019