Keywords
drug discovery
fragment based drug discovery
hit optimization
ligand efficiency
molecular modeling
scoring function
Abstract
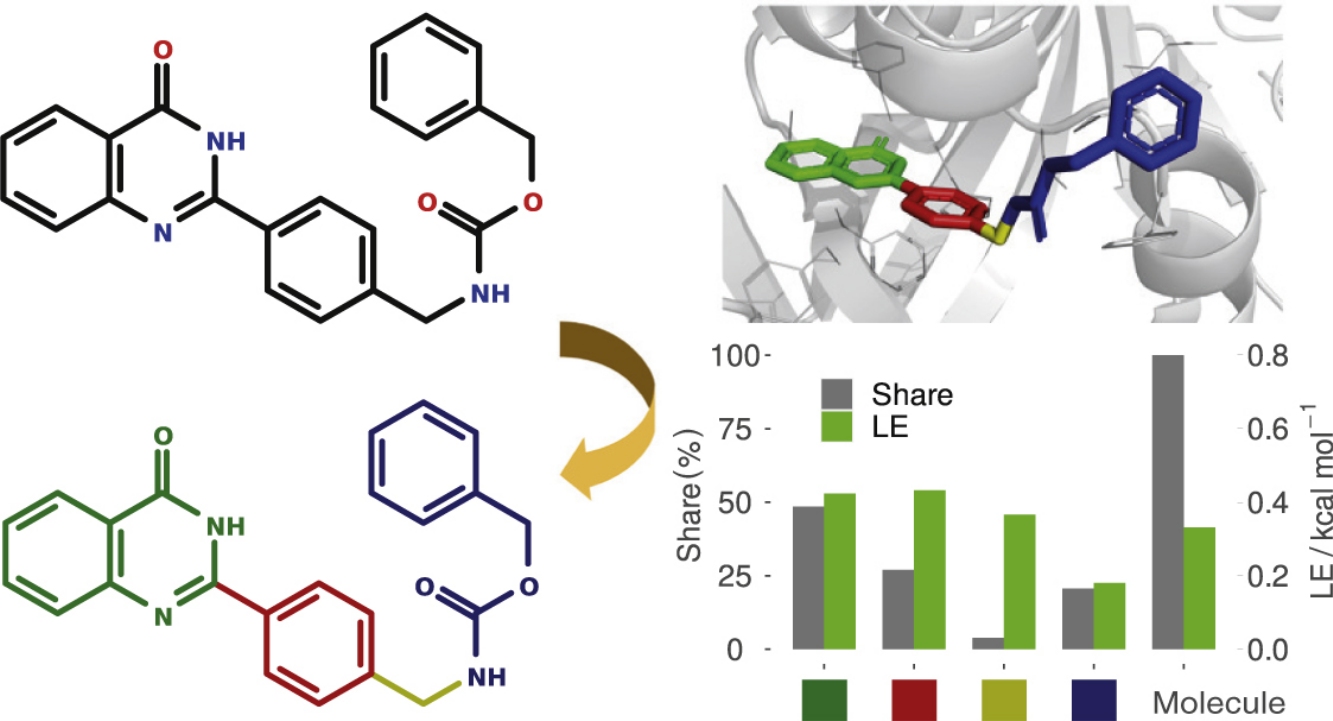
Contributions of different fragments of a ligand into the binding/activity to a specified target are of importance to guide hit-to-lead drug discovery, and fragment based drug discovery (FBDD) approach has proven to be quite fruitful. However, the experimental means of FBDD are generally not affordable to many researchers working in the drug discovery field, especially to small medicinal chemistry groups at universities. To partially solve this problem, we propose a Reversed Fragment Based Drug Discovery (R-FBDD) approach in which the contributions of fragments of a molecule are estimated using scoring functions in order to detect whether a fragment is a ‘binding anchor’ or a ballast, thus guiding further development.
References
1.
10.1016/j.mencom.2021.04.004_bib0005
Congreve
Annu. Rep. Med. Chem.,
2007
2.

Joseph-McCarthy D., Campbell A.J., Kern G., Moustakas D.
Journal of Chemical Information and Modeling,
2014
3.
10.1016/j.mencom.2021.04.004_bib0015
Rydzewski
Real World Drug Discovery: A Chemist's Guide to Biotech and Pharmaceutical Research,
2010
4.

Whitty A.
Future Medicinal Chemistry,
2011
5.

Liu T., Naderi M., Alvin C., Mukhopadhyay S., Brylinski M.
Journal of Chemical Information and Modeling,
2017
6.

Hopkins A.L., Keserü G.M., Leeson P.D., Rees D.C., Reynolds C.H.
Nature Reviews Drug Discovery,
2014
7.

Trott O., Olson A.J.
Journal of Computational Chemistry,
2009
8.

Morris G.M., Huey R., Lindstrom W., Sanner M.F., Belew R.K., Goodsell D.S., Olson A.J.
Journal of Computational Chemistry,
2009
9.

O'Boyle N.M., Banck M., James C.A., Morley C., Vandermeersch T., Hutchison G.R.
Journal of Cheminformatics,
2011
10.

Nathubhai A., Haikarainen T., Hayward P.C., Muñoz-Descalzo S., Thompson A.S., Lloyd M.D., Lehtiö L., Threadgill M.D.
European Journal of Medicinal Chemistry,
2016
11.

Zaryanova E.V., Lozinskaya N.A., Beznos O.V., Volkova M.S., Chesnokova N.B., Zefirov N.S.
Bioorganic and Medicinal Chemistry Letters,
2017