Keywords
disulfiram
main protease Mpro
molecular docking
molecular dynamics
neratinib
SARS-CoV-2
Abstract
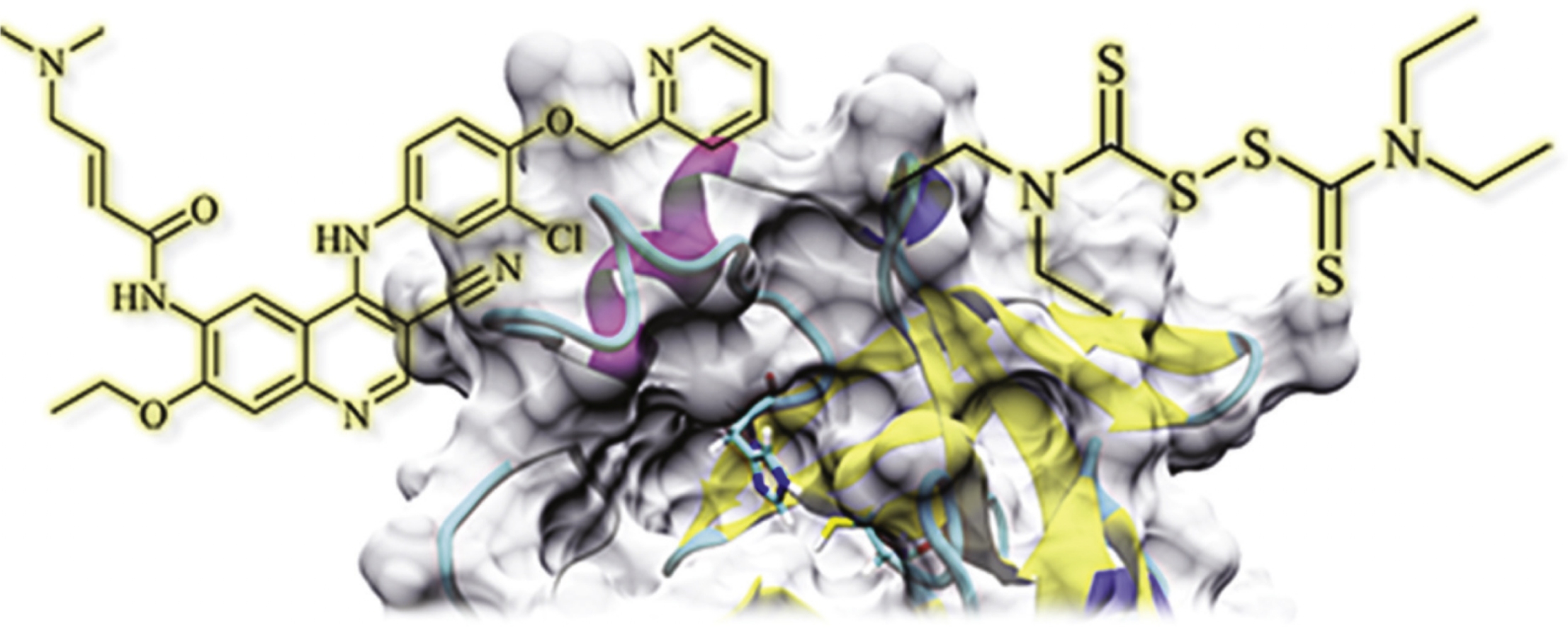
Identification of disulfiram and neratinib as putative covalent inhibitors of SARS-CoV-2 virus main protease Mpro by a combination of ‘on-top docking’ procedure, expert evaluation of potential hits and molecular dynamics is reported herein. This finding shows the importance of further development of virtual screening add-ons.
References
1.

Novikov F.N., Stroylov V.S., Svitanko I.V., Nebolsin V.E.
Russian Chemical Reviews,
2020
2.
10.1016/j.mencom.2020.07.004_bib0010
Jin
Nature,
2020
3.

Cao B., Wang Y., Wen D., Liu W., Wang J., Fan G., Ruan L., Song B., Cai Y., Wei M., Li X., Xia J., Chen N., Xiang J., Yu T., et. al.
New England Journal of Medicine,
2020
4.

Kandeel M., Al-Nazawi M.
Life Sciences,
2020
5.

Douguet D.
ACS Medicinal Chemistry Letters,
2018
6.

Stroganov O.V., Novikov F.N., Stroylov V.S., Kulkov V., Chilov G.G.
Journal of Chemical Information and Modeling,
2008
7.
M. Medvedev, O. Stroganov, A. Dmitrienko, M. Panova, I. Svitanko, F. Novikov and G. Chilov, J. Chem. Inf. Model., in press.
8.

Albulescu L., Hale M.S., Ainsworth S., Alsolaiss J., Crittenden E., Calvete J.J., Evans C., Wilkinson M.C., Harrison R.A., Kool J., Casewell N.R.
Science Translational Medicine,
2020
9.
10.1016/j.mencom.2020.07.004_bib0045
Lobo-Galo
J. Biomol. Struct. Dyn.,
2020
10.

Abraham M.J., Murtola T., Schulz R., Páll S., Smith J.C., Hess B., Lindahl E.
SoftwareX,
2015
11.

Dodda L.S., Cabeza de Vaca I., Tirado-Rives J., Jorgensen W.L.
Nucleic Acids Research,
2017
12.
10.1016/j.mencom.2020.07.004_bib0060
Xu
ChemRxiv,
2020
13.

Lin M., Moses D.C., Hsieh C., Cheng S., Chen Y., Sun C., Chou C.
Antiviral Research,
2018
14.
10.1016/j.mencom.2020.07.004_bib0070
Horowitz
Respir. Med. Case Rep.,
2020
15.
10.1016/j.mencom.2020.07.004_bib0075
Polonikov
ACS Infect. Dis.,
2020
16.

Chuck C., Chen C., Ke Z., Chi-Cheong Wan D., Chow H., Wong K.
European Journal of Medicinal Chemistry,
2013
17.

Stroganov O.V., Novikov F.N., Medvedev M.G., Dmitrienko A.O., Gerasimov I., Svitanko I.V., Chilov G.G.
Journal of Computer-Aided Molecular Design,
2020
18.
Khrenova M.G., Tsirelson V.G.
Mendeleev Communications,
2019