Abstract
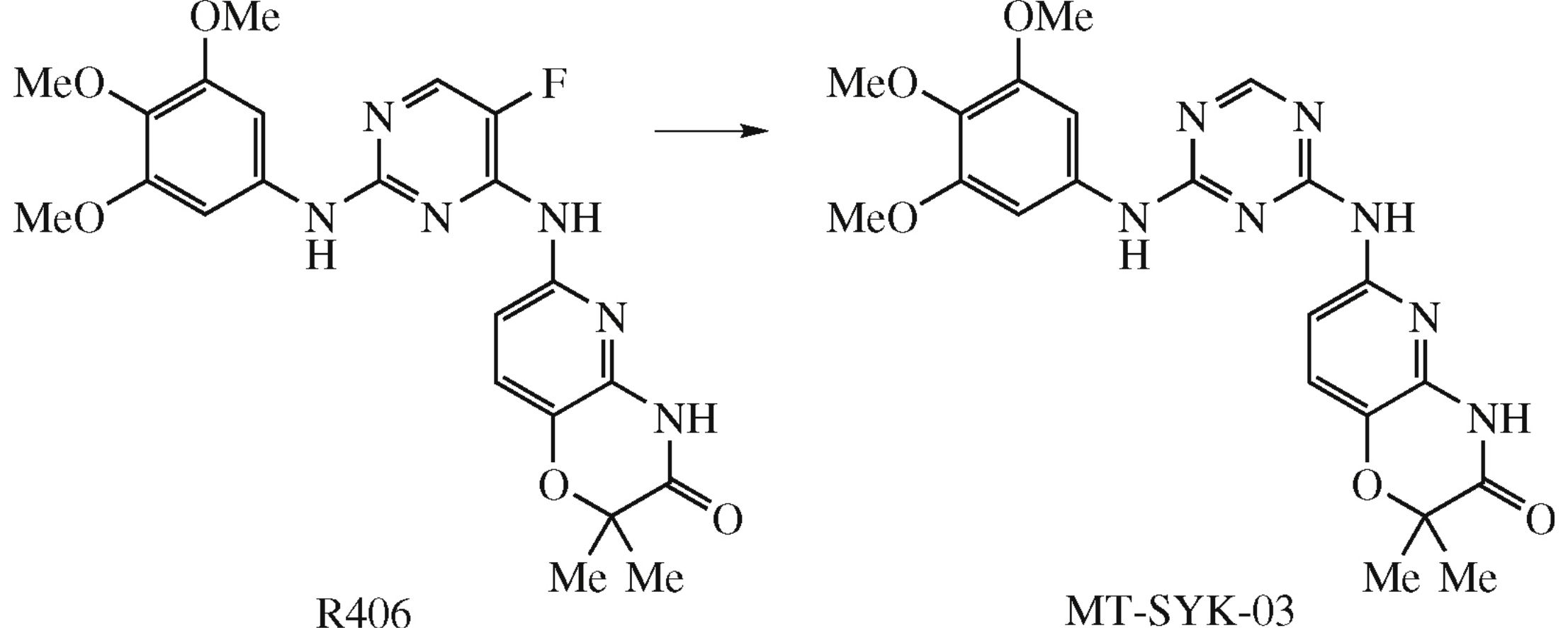
Free energy perturbation (FEP)-based molecular modeling simulation of 5-fluoropyrimidine and 1,3,5-triazine derivatives followed by their synthesis and experimental evaluation have been carried out to estimate kinase selectivity profile. 5-Fluoropyrimidine derivatives show similar binding affinity for c-Src, Btk and Jak1 kinases, while 1,3,5-triazine derivatives demonstrate c-Src kinase selectivity.
References
1.

Rolf M.G., Curwen J.O., Veldman‐Jones M., Eberlein C., Wang J., Harmer A., Hellawell C.J., Braddock M.
Pharmacology Research and Perspectives,
2015
2.
Rakitina T.V., Zeifman A.A., Titov I.Y., Svitanko I.V., Lipkin A.V., Stroylov V.S., Stroganov O.V., Novikov F.N., Chilov G.G.
Mendeleev Communications,
2012
3.

Zeifman A.A., Stroylov V.V., Novikov F.N., Stroganov O.V., Kulkov V., Chilov G.G.
Journal of Chemical Theory and Computation,
2013
4.

Cowan-Jacob S.W., Fendrich G., Manley P.W., Jahnke W., Fabbro D., Liebetanz J., Meyer T.
Structure,
2005
5.

Kuglstatter A., Wong A., Tsing S., Lee S.W., Lou Y., Villaseñor A.G., Bradshaw J.M., Shaw D., Barnett J.W., Browner M.F.
Protein Science,
2011
6.

Vasbinder M.M., Alimzhanov M., Augustin M., Bebernitz G., Bell K., Chuaqui C., Deegan T., Ferguson A.D., Goodwin K., Huszar D., Kawatkar A., Kawatkar S., Read J., Shi J., Steinbacher S., et. al.
Bioorganic and Medicinal Chemistry Letters,
2016
7.

Stroganov O.V., Novikov F.N., Zeifman A.A., Stroylov V.S., Chilov G.G.
Proteins: Structure, Function and Genetics,
2011
8.

Stroganov O.V., Novikov F.N., Stroylov V.S., Kulkov V., Chilov G.G.
Journal of Chemical Information and Modeling,
2008
9.
Prokhorov E.I., Bekker A.V., Perevoznikov A.V., Kumskov M.I., Svitanko I.V.
Mendeleev Communications,
2015
10.

Van Der Spoel D., Lindahl E., Hess B., Groenhof G., Mark A.E., Berendsen H.J.
Journal of Computational Chemistry,
2005
11.

Jorgensen W.L., Maxwell D.S., Tirado-Rives J.
Journal of the American Chemical Society,
1996
12.

Sousa da Silva A.W., Vranken W.F.
BMC Research Notes,
2012
13.

Wang J., Wang W., Kollman P.A., Case D.A.
Journal of Molecular Graphics and Modelling,
2006
14.
Stroylov V.S., Rakitina T.V., Novikov F.N., Stroganov O.V., Chilova G.G., Lipkin A.V.
Mendeleev Communications,
2010
15.

Mian A.A., Rafiei A., Haberbosch I., Zeifman A., Titov I., Stroylov V., Metodieva A., Stroganov O., Novikov F., Brill B., Chilov G., Hoelzer D., Ottmann O.G., Ruthardt M.
Leukemia,
2014
16.

Kemble D.J., Wang Y., Sun G.
Biochemistry,
2006