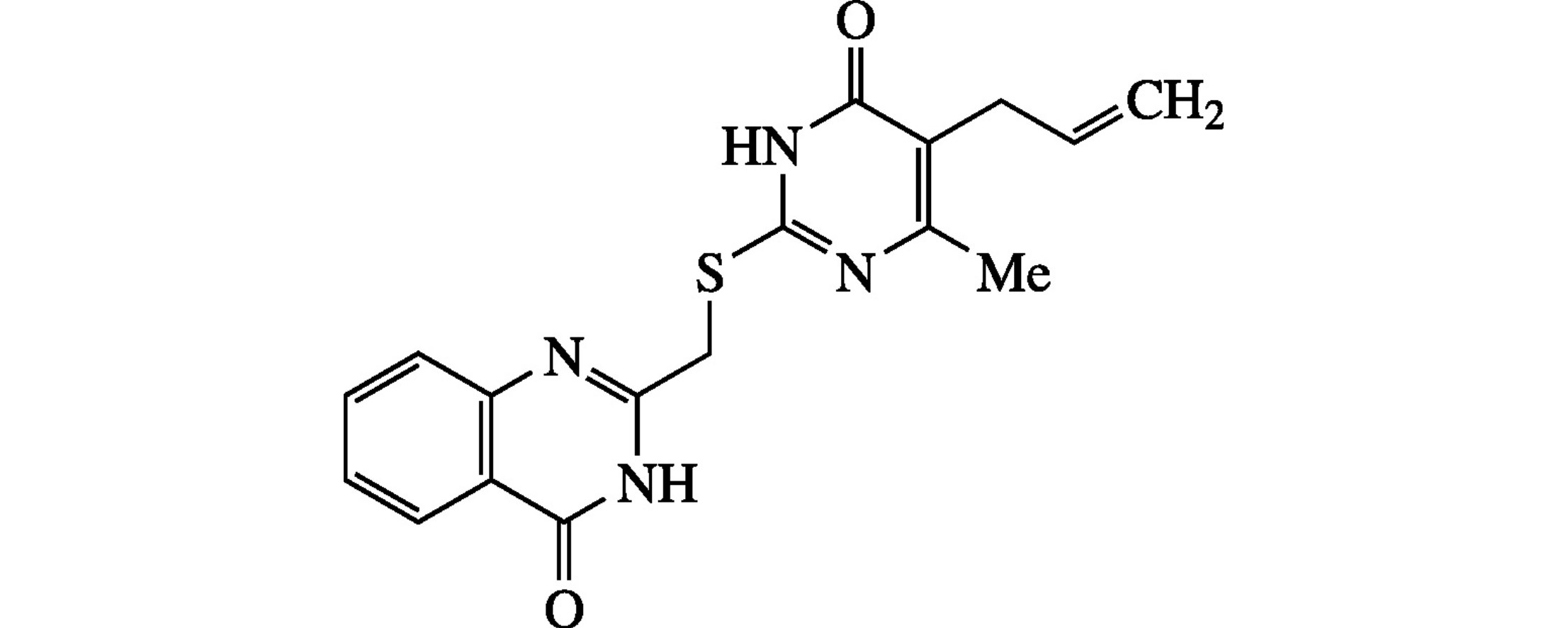
Abstract
3D-QSAR and molecular docking were applied to predict the inhibitory activity of 196 compounds towards poly(ADP-riboso) polymerase-1 (PARP). A proportion of experimentally active ligands was higher among compounds with good rankings from both methods (57%) compared to compounds scored as inactive by at least one method (40% for docking-active, QSAR-inactive compounds).
References
1.
Stahura F., Bajorath J.
Combinatorial Chemistry and High Throughput Screening,
2004
2.

Sousa S.F., Ribeiro A.J., Coimbra J.T., Neves R.P., Martins S.A., Moorthy N.S., Fernandes P.A., Ramos M.J.
Current Medicinal Chemistry,
2013
3.
Zefirov N.S., Palyulin V.A.
Mendeleev Communications,
2010
4.
Gazieva G.A., Vikharev Y.B., Anikina L.V., Karpova T.B., Kravchenko A.N., Permyakov E.A., Svitanko I.V.
Mendeleev Communications,
2013
5.
Romashov L.V., Zeifman A.A., Zakharenko A.L., Novikov F.N., Stroilov V.S., Stroganov O.V., Chilov G.G., Khodyreva S.N., Lavrik O.I., Titov I.Y., Svitan’ko I.V.
Mendeleev Communications,
2012
6.

d'AMOURS D., DESNOYERS S., d'SILVA I., POIRIER G.G.
Biochemical Journal,
1999
7.

Stroganov O.V., Novikov F.N., Stroylov V.S., Kulkov V., Chilov G.G.
Journal of Chemical Information and Modeling,
2008
8.

Stroganov O.V., Novikov F.N., Zeifman A.A., Stroylov V.S., Chilov G.G.
Proteins: Structure, Function and Genetics,
2011
9.

Novikov F.N., Zeifman A.A., Stroganov O.V., Stroylov V.S., Kulkov V., Chilov G.G.
Journal of Chemical Information and Modeling,
2011
10.

Novikov F.N., Stroylov V.S., Stroganov O.V., Chilov G.G.
Journal of Molecular Modeling,
2009
11.
F. N. Novikov, PhD Thesis, Moscow, 2011.
12.

Prokhorov E.I.
Moscow University Mathematics Bulletin,
2013
13.

Bekker A.V., Suleymanov A.A., Apryshko G.N., Kumskov M.I.
Pattern Recognition and Image Analysis,
2011
14.

Perevoznikov A.V., Shestov A.M., Permyakov E.A., Kumskov M.I.
Pattern Recognition and Image Analysis,
2011
15.
10.1016/j.mencom.2015.05.019_bib0070
Kumskov
Dokl. Akad. Nauk,
1994